Gene Ontology
Gene ontology was developed as a way to unify gene and gene product attributes across all species. This was done by creating a unified vocabulary of attributes, annotating genes and gene products, and making this data easily accessible. Gene ontology is comprised of three categories: biological process, cellular component, and molecular function. A biological process of a gene is the biological goal that the gene aids in completing. The cellular component is the location where the gene is active in the cell. The molecular function is the biochemical activity that the gene performs [1]. All of these are important in identifying the role a certain gene plays in a cell. By identifying this purpose, it can be easier to understand how diseases function and more importantly how treatments can be developed for them. The program QuickGo can be used to find out the ontology of different genes. The ontology of SMPD1 is listed below [2] [3].
SMPD1 Ontology
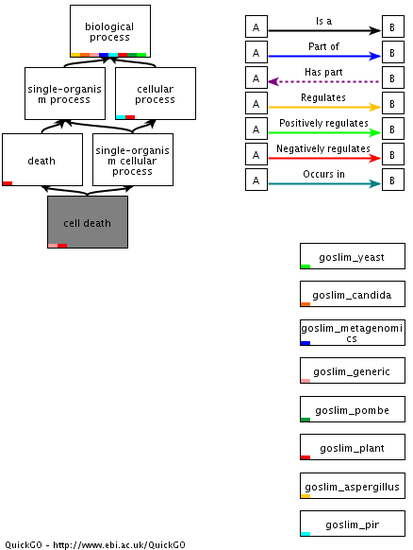
Biological Process:Sphingomyelin catabolic & metabolic processes
Cell death Nervous system development |
Cellular Component:Extracellular space
Lysosomal lumen Lamellar body |
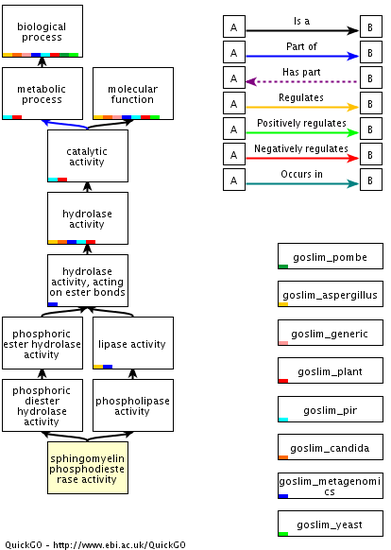
Molecular Function:Sphingomyelin phosphodiesterase activity
Hydrolase activity Acting on glycosyl bonds |
Analysis and Discussion
The different roles that SMPD1 plays in the cell are very important to proper functioning. When there is a mutation in the gene, such as in Niemann-Pick patients, these functions are inhibited or altered. The biological function of cell death is logical because ceramide is needed to trigger cell death, and there is a lack of ceramide in patients with Niemann-Pick. SMPD1 is found in lysosomes, which is why one of the cellular components is lysosomal lumen. The molecular function of sphingomyelin phosphodiesterase activity is also inhibited in Niemann-Pick. This molecular function is illustrated in the figure on the right above.