What is Phylogeny?
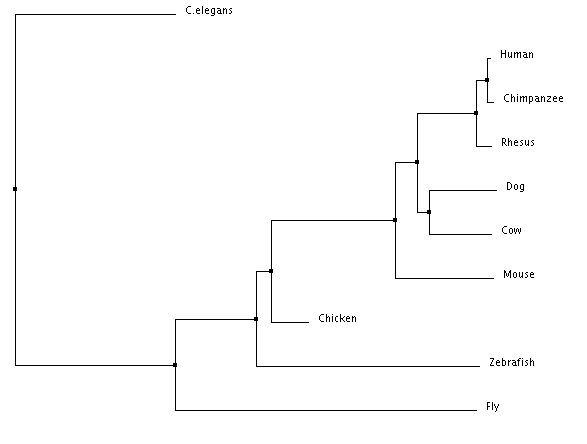
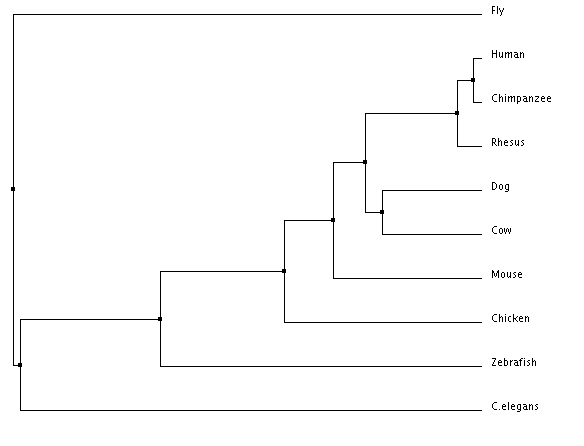
Phylogeny is the physical mapping of protein sequences based off of the homology of the sequences. Phylogenetic trees can show the evolutionary history of organisms. The phylogenetic trees were created using ClustalOmega, which allows for a multiple sequence alignment. The protein sequences for nine organisms were inputed into the program, which then aligned them into the trees below. The neighbor joining tree aims to make the shortest tree possible, and the average distance tree puts organisms that are the most closely related branching from the same node.
Phylogenetic Trees
Analysis and Discussion
The neighbor joining percent identity tree shows C. elegans as being the outgroup, whereas the average distance percent identity tree shows flies as being the outgroup. The trees show where in time the species diverge. The human and the chimpanzee have the closest related protein sequences, which is why they are in a sub-group together. They diverged from a common ancestor more recently than the dog and the cow did, who are also in a sub-group together. The trees are very similar to the homology showing that vertebrates protein sequences are more closely related to humans than invertebrates.
References
ClustalOmega
ClustalOmega